Automated Testing for Microservices
How Audiogum's testing strategies help us maintain a high pace of change.
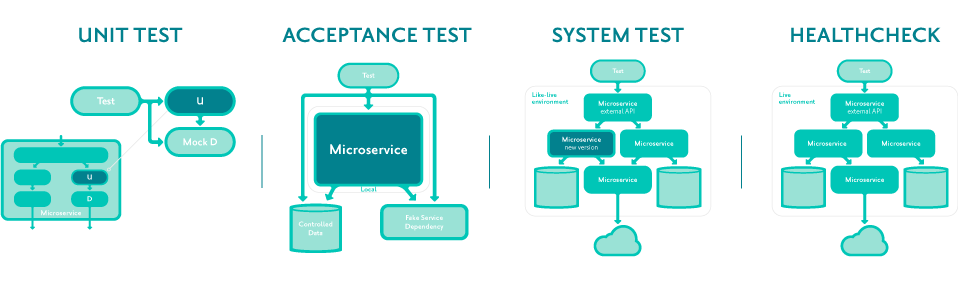
Overview
At Audiogum, to achieve both high availability and rapid pace of change, we rely on repeatable, fully automated testing of our services.
Unit tests, acceptance tests, component tests, integration tests, system tests, end-to-end tests... These are just a few of the commonly used terms to describe different flavours of testing, but there is ambiguity in what these things mean and how they relate, as well as endless potential implementations.
In this post I aim to describe what we Audiogum engineers agree as reasonable definitions for the kinds of tests we think are important in the context of our microservices architecture, as described by Steve previously in the platform overview post. For each of these I describe the motivations and implications of these definitions and some strategies we have adopted at Audiogum that suit our context.
We have other types of testing such as performance and load testing that are important too but I will leave these out of scope and focus for now on the testing of service behaviour.
Motivation: Tests Support Change
We test because we need to confirm as best we can that what we create behaves as intended. Furthermore, testing helps us to prevent breaking things when we change them.
All too often though, tests can become barriers rather than enablers for change. They take time to create, to run, and to maintain. Our motivation has been to avoid some of the problems we've each encountered elsewhere in the past, including for example:
- Duplication of tests across multiple levels of testing -> costs time
- Inconsistency of approach causing confusion for new engineers on a project -> costs time
- Tests that are hard to understand therefore difficult to maintain -> costs time
- Tests that are hard to run due to fragile or hard to set up dependencies -> costs time
- Tests that are ignored, pointless or incorrectly modified (often due to above) -> risk of regression and escaped bugs
Unit Tests
Unit Tests: Audiogum definition
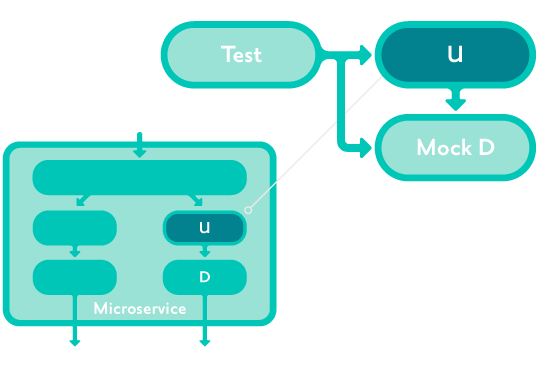
Unit tests are automated tests of small parts of a service. For example in our Clojure microservices, a function or set of functions in a namespace.
Unit tests avoid external dependencies (no I/O) by providing mocked implementations of dependencies wherever necessary.
Unit tests check the current internal implementation NOT overall system behaviour.
Unit Tests: Motivation
Whether taking the "test first" approach, writing tests afterwards or somewhere in between, unit tests should support the development of relatively detailed logic, independently of other parts of the system.
Programming languages like Clojure with its REPL allow us to interact directly with running code and data, testing as we write it. This is a large part of the productivity benefit we perceive in Clojure. Becoming comfortable with a REPL-based approach to programming can mean that unit tests no longer seem essential to the developer to feel confident that the details work as they intend as they create the code. However, unit tests create a repeatable record of such testing and so can be valuable for both the initial implementer and any future developers maintaining and extending the code later. They help prove that new changes haven't broken previous behaviour unintentionally and they convey understanding of how the implementation is intended to work.
Unit Tests: Implications
Unit tests must be fast to run and easy to maintain so they are not blockers for change.
Unit tests are optional and disposable because they are useful only for internal details. We accept that refactoring breaks existing tests and they may need replacement or modification. Some services actually have very little internal logic and are mainly concerned with I/O and wiring other things together. In these cases it's OK to have few or no unit tests at all. Other services perform some algorithmic process or complex transformation logic internally. It's in these cases where unit testing has value as described above.
Therefore while code coverage can be an interesting indicator of test health, we do NOT consider it a goal in itself. 100% unit test code coverage for an entire microservice is impossible by our definition.
We certainly cannot rely on unit tests alone to check overall service behaviour - hence the other levels below!
Unit Tests: Audiogum implementation
Audiogum's microservice unit tests are implemented in Clojure and run in-process with no network I/O. The tests run locally on demand and automatically as part of the CI job for the project.
Dependencies of the code under test, if any, are stubbed or mocked.
For readers interested in Clojure, here's a Clojure-specific interlude on dependencies:
Past experience using testing frameworks like Midje has caused a tendancy to redefine hard dependencies (e.g. using against-background or provided). One can achieve similar "under the hood" fiddling using something like with-redefs.
At Audiogum we believe that to avoid this style leads to better code structure, specifically separation of concerns. We prefer as much of our code to be pure as possible and avoid having hidden dependencies on non-deterministic or external stuff. Pure code is predictable and therefore testable!
Using clojure.test and avoiding function redefinition encourages us to make our code as pure as possible and clearly separate concerns by passing function dependencies and creating abstractions such as protocols where appropriate.
Unit testing such code then involves creating test implementations of the dependencies and passing them into the code under test. There is often nothing more to this than defining stub functions, but there are libraries such as Shrubbery we sometimes use to support more advanced mocking, in this case when using protocols specifically. Our approach to unit testing prompted the development of the Picomock library, which provides some simple convenience tools for testing with dependencies and does not depend on protocols.
Acceptance Tests
Acceptance Tests: Audiogum definition
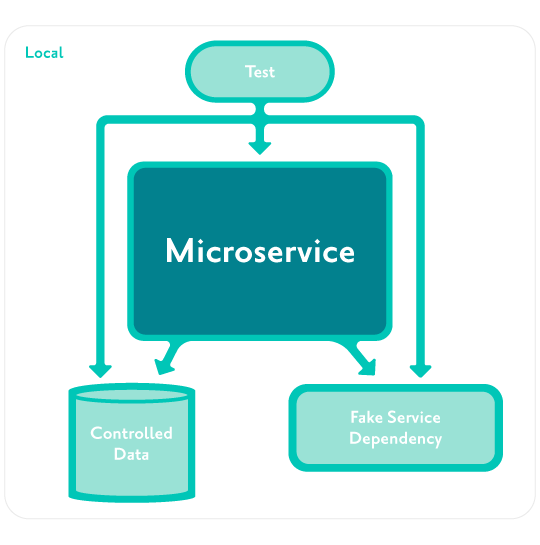
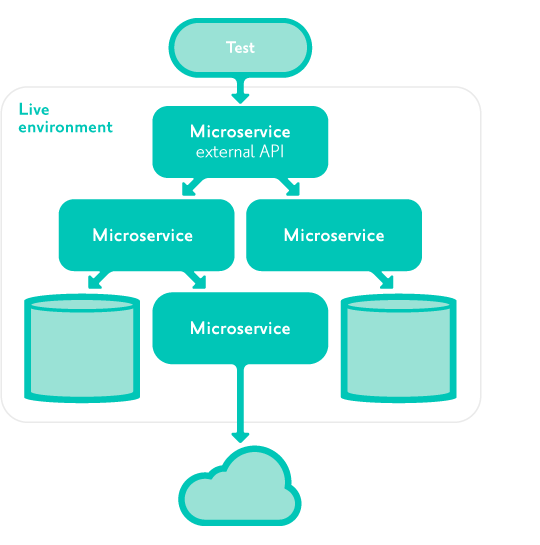
Acceptance tests are automated tests run against the external API of an instance of a microservice isolated from other environments.
The tests do not interact directly with the internals of the service.
Dependencies of the service are also isolated, either fixed or controlled by the tests.
Acceptance Tests: Motivation
Acceptance tests prove the correct external behaviour of a microservice, including its I/O boundaries, given specific known behaviours of its dependencies. This supports change and internal refactoring: we should be confident to change internal implementation as long as acceptance tests still pass - in contrast to disposable unit tests above!
Isolation of these tests and their dependencies is essential to allow developers to do their work independently, and to allow such tests to run reliably: there should be nothing outside the control of the project that can make them fail.
Acceptance Tests: Implications
We must ensure coverage of expected behaviour of each service is as complete as possible in its acceptance tests in order to prevent accidental breakage when adding new features or refactoring.
However, acceptance tests with fake dependencies as described are not enough alone to support continuous deployment of microservices within the larger system. The tests themselves may make incorrect or incomplete assumptions about dependencies. Hence the need for system tests as described later!
Acceptance Tests: Audiogum implementation
Our acceptance tests consist of Clojure code running against the API of a microservice (generally a REST API, e.g. using clj-http.client).
The service is hosted locally but not in the same process as the tests. This supports the idea that the tests don't interact with the internals by preventing us "accidentally" mocking out internal parts.
Like unit tests, the acceptance tests can be run locally on demand and form part of the CI job.
External dependencies of the microservice are substituted for local surrogates controlled by the test. This is achieved by configuration of endpoints, e.g. using environment variables to supply URLs, NOT by changing the protocols or internal implementation.
For example:
- Dependency on DynamoDb: we use Amazon's local DynamoDb, started and populated by the test
- Dependency on another microservice or external REST API: we use rest driver via rest-cljer to create fake responses and check for expected requests
System Tests
System Tests: Audiogum definition
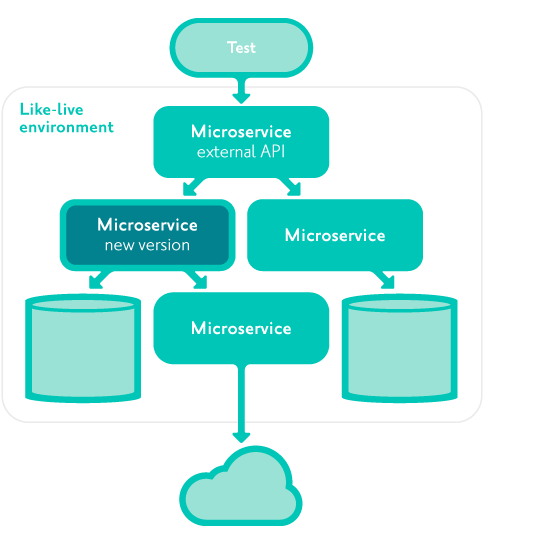
System tests are automated tests interacting only with the external client-facing APIs of the system as a whole, not with any internal microservice APIs or databases.
They are run in a 'like-live' environment where all dependencies are identical versions to production.
They are run whenever a change is being prepared for production, prior to deployment to production.
System Tests: Motivation
System tests prove the correct behaviour of the entire system from the client user's point of view.
They support continuous delivery by determining that a new version of a microservice will behave correctly in the context of the currently live clients and dependencies. Since they use real dependencies they can catch certain kinds of problems that acceptance tests might not.
They also support re-engineering within the microservices system, in the same way as each service's acceptance tests support internal refactoring: for example we can change how the microservices interact with each other as long as we don't break the behaviour of the external API.
System Tests: Implications
We should ensure all important user-facing features are represented and all dependencies are exercised by system tests. However we do not need to cover all detailed behaviours in system tests - to do so would cause too much duplication of acceptance tests and therefore increased barrier to change.
Since system tests run in a 'like-live' environment rather than actual production, even with perfect versioning and environment isolation they cannot prove that the production system is still working. That's where healthchecks come in, as described later!
System Tests: Audiogum implementation
In our definition we stated "dependencies are identical versions to production" and "run whenever a change is being prepared ... prior to deployment to production". The intention of course is to prevent other changes from affecting the test and giving us the wrong result. Strictly speaking this means we need complete isolation of the environment in which we are testing, and to change only one thing (microservice for example) at a time.
This presents a challenge: How can we provide this kind of isolation without blocking each other's independent work? We choose a microservices architecture specifically because it supports fast independent change!
At Audiogum we have considered various potential solutions, including:
- The 'delivery pipeline' approach: Assuming a shared environment with all versions as per production, we automate a process such that we update only one microservice then run the system tests. Deploy to production if it succeeds. While we do all of this, block any other changes to the testing environment.
- The 'disposable environments' approach: Create isolated environments on demand, potentially simultaneously, with all other versions as per production, in which to run the system tests for any change. Deploy that one change to production if it succeeds.
The pipeline approach is a well understood and common solution to this kind of problem, however it has the potential to cause a bottleneck. We can imagine reducing the run time as much as possible to mitigate this, but there will still be blockages, particularly when tests fail and we want to fix forward.
The disposable environments approach solves this by allowing anyone to run system tests with their own change independently. We trade this at the cost of potential incorrectness: there's nothing to prevent another change happening such that the production dependency versions are not the same as when we created the test environment. This may be a risk worth taking for the sake of independence and speed, but we must also consider the cost of automating the creation of the environment: not trivial in the case of a distributed system composed of many microservices, data stores etc.
At the time of writing, we have neither a strict pipeline nor disposable environments, but rather a compromise that involves trust and collaboration in favour of complexity and strictness.
We use a single 'staging' environment (involving among other pieces a Kubernetes cluster hosted in AWS), identical to production but isolated from it. Engineers can deploy updates to any service at any time. The system tests, implemented again in Clojure, can be run against the staging environment on demand but also run on a regular schedule. The general rule is that each developer is resposible for checking that their change, having passed acceptance tests, also does not break the system tests after deploying to staging, before promoting the change to production.
While we are a small team collaborating continuously, this works well for us. We anticipate that as we expand the team, or increase the size and complexity of the overall system, we will take the next steps in order to maintain the pace of independent change at larger scale. A disposable environments approach is feasible for us since our infrastructure setup is already fully automated using Terraform.
Healthcheck Tests
Healthcheck Tests: Audiogum definition
Healthcheck tests are automated tests interacting with the production system exactly as would a real client. They are run on a frequent scheduled basis, irrespective of changes.
Healthcheck Tests: Motivation
Healthchecks alert us when, despite the testing performed before deployment to production, some change breaks an important function of the system. This allows us to roll back or fix such changes quickly without relying on customers reporting a problem.
They can also alert us of breaks in functional availability that might not be picked up by system metric monitoring or logging, for example due to infrastructure changes, or 3rd party dependencies changing.
Healthcheck Tests: Implications
Since they run in production, it may not always be possible or appropriate to test every detailed feature, although as per system tests they should aim to exercise every dependency.
Since we want fast feedback, sanity tests should be as quick to run as possible.
Healthcheck Tests: Audiogum implementation
Audiogum's healthcheck tests work essentially the same as the system tests, except they run against the production environment rather than staging. In fact we run identical test code, but we flag which tests are appropriate for production.
Just like system tests, we run the healthchecks on demand but also on a regular schedule and we have automated alerting when something fails. More about how this fits into our availability monitoring and alerting strategy can be found in Deon's post: Audiogum Service Status.
Conclusion
“Everything should be as simple as it can be, but not simpler”
It may seem that implementing an automated testing strategy with multiple levels must be arduous, but actually with pragmatism at each level, techniques like these can enable teams to work on large and necessarily complex systems with greater speed and safety.
The most important thing is that when problems do occur, we can find and fix them as quickly as possible. We do this with the help of trust and tools that facilitate changes, not block them.
